Chemical Properties:
Octanol-water partition coefficient (log P) : The partition coefficient, abbreviated P, is defined as a particular ratio of the concentrations of a solute between the two solvents (a biphase of liquid phases), specifically for un-ionized solutes, and the logarithm of the ratio is thus log P [1].
Molecular weight (MW): Molecular weight also called molecular mass, mass of a molecule of a substance, based on 12 as the atomic weight of carbon-12. It is calculated in practice by summing the atomic weights of the atoms making up the substance’s molecular formula.
Hydrogen bond acceptor (HBA): The atom, ion, or molecule component of a hydrogen bond which does not supply the bridging (shared) hydrogen atom.
Hydrogen bond donor (HBD): A bond or molecule that supplies the hydrogen atom of a hydrogen bond.
Rotatable bonds (RotB): The number of rotatable bonds is the number of bonds which allow free rotation around themselves.
Topological polar surface area (TPSA) : The topological polar surface area (TPSA) of a molecule is defined as the surface sum over all polar atoms or molecules, primarily oxygen and nitrogen, also including their attached hydrogen atoms.
Quantitative Estimation of Druglikeness (QED) : QED provides a quantitative metric for assessing drug-likeness. The provided values can range between zero (all properties unfavorable) and one (all properties favorable). The functions are based on the underlying distribution data of drug properties [2].
Synthetic Accessibility score (SAscore): The method for estimation of the synthetic accessibility score (SAscore) described here is based on a combination of fragment contributions and a complexity penalty [3].
- Wildman, S.A. and Crippen, G.M. (1999) Prediction of Physicochemical Parameters by Atomic Contributions. Journal of Chemical Information and Computer Sciences, 39, 868-873.
- Bickerton, G.R., Paolini, G.V., Besnard, J., Muresan, S. and Hopkins, A.L. (2012) Quantifying the chemical beauty of drugs. Nat Chem, 4, 90-98.
- Ertl, P. and Schuffenhauer, A. (2009) Estimation of synthetic accessibility score of drug-like molecules based on molecular complexity and fragment contributions. Journal of Cheminformatics, 1, 8.
How To:
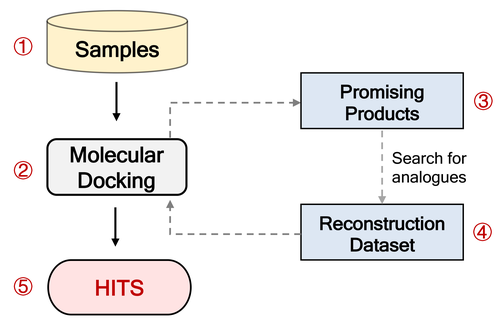
Step 1. Sample(s) preparation. One first needs to download the samples from DrugSpaceX. There are two ways here, users can download analogs of the drug of interest, or download representative samples from DrugSpaceX (https://drugspacex.simm.ac.cn/download/) for subsequent virtual screening. Representative samples are recommended to download the drug set or 1st transformed S sample, both of which provide corresponding structure smiles (.smi), 2D and 3D structures (.sdf) for docking.
Step 2. When the ligands are ready, the compounds are docked into the target of interest. Our goal is to find synthesizable and novel chemical structure with a desirable biological effect. Generally, we choose the molecules with the highest docking score.
Step 3. We use the docking method to find promising molecules based on the docking scoring results. The next step, we analyze and exactly search the structure. On the one hand, other than chemical properties and transformation instructions, DrugSpaceX could also locate the position of transformation, which enables medicinal chemists easily systematically integrate strategy planning and protection design purposes. On the other hand, we can download the drug analogues to reconstruct dataset for molecular docking again.
Step 4. Based on the similarity search results, the dataset can be reconstructed to further discover promising molecules.
Step 5. Find Hits through the above operation.